The MsBackendPy allows to access MS data stored as matchms.Spectrum
or spectrum_utils.spectrum.MsmsSpectrum objects from the
matchms respectively
spectrum_utils Python
library directly from R. The MS data (peaks data or spectra variables) are
translated on-the-fly when accessed. Thus, the MsBackendPy allows a
seamless integration of Python MS data structures into Spectra::Spectra()
based analysis workflows.
The MsBackendPy object supports replacing values for peaks variables
(m/z and intensity) and adding/replacing or removing spectra variables.
The changes are immediately translated and written back to the Python
variable.
See the descripion of the backendInitialize() method below for creation
and initialization of objects from this class. Also, the setBackend()
method for Spectra::Spectra() objects internally uses
backendInitialize(), thus the same parameters can (and have) to be passed
if the backend of a Spectra object is changed to MsBackendPy using
the setBackend() method. Special care should also be given to parameter
spectraVariableMapping, that defines which spectra variables should be
considered/translated and how their names should or have to be converted
between R and Python. See the description for backendInitialize() and the
package vignette for details and examples.
Usage
# S4 method for class 'MsBackendPy'
backendInitialize(
object,
pythonVariableName = character(),
spectraVariableMapping = defaultSpectraVariableMapping(),
pythonLibrary = c("matchms", "spectrum_utils"),
...,
data
)
# S4 method for class 'MsBackendPy'
length(x)
# S4 method for class 'MsBackendPy'
spectraVariables(object)
# S4 method for class 'MsBackendPy'
spectraData(object, columns = spectraVariables(object), drop = FALSE)
# S4 method for class 'MsBackendPy'
spectraData(object) <- value
# S4 method for class 'MsBackendPy'
peaksData(object, columns = c("mz", "intensity"), drop = FALSE)
# S4 method for class 'MsBackendPy'
peaksData(object) <- value
# S4 method for class 'MsBackendPy'
x$name
# S4 method for class 'MsBackendPy'
x$name <- value
# S4 method for class 'MsBackendPy'
intensity(object) <- value
# S4 method for class 'MsBackendPy'
mz(object) <- value
# S4 method for class 'MsBackendPy'
spectraNames(object) <- value
# S4 method for class 'MsBackendPy'
spectraVariableMapping(object) <- value
# S4 method for class 'MsBackendPy'
spectraVariableMapping(object, value)
reindex(object)
# S4 method for class 'Spectra,MsBackendPy'
setBackend(
object,
backend,
pythonVariableName = character(),
spectraVariableMapping = defaultSpectraVariableMapping(),
pythonLibrary = c("matchms", "spectrum_utils"),
applyProcessing = TRUE,
...
)Arguments
- object
A
MsBackendPyobject.- pythonVariableName
For
backendInitialize()andsetBackend():character(1)with the name of the variable/Python attribute that contains the list ofmatchms.Spectrumobjects with the MS data.- spectraVariableMapping
For
backendInitialize()andsetBackend(): namedcharacterwith the mapping between spectra variable names and (matchms.Spectrum) metadata names. SeedefaultSpectraVariableMapping(), and the description of thebackendInitialize()function forMsBackendPyfor more information and details. Note that forsetBackend()only spectra variables defined by this parameter are transferred to Python.- pythonLibrary
For
backendInitialize()andsetBackend():character(1)specifying the Python library used to represent the MS data in Python. Can be eitherpythonLibrary = "matchms"(the default) orpythonLibrary = "spectrum_utils".- ...
Additional parameters.
- data
For
backendInitialize():DataFramewith the full MS data (peaks data and spectra data) such as extracted with theSpectra::spectraData()method on anotherMsBackendinstance. Importantly, theDataFramemust have columns"mz"and"intensity"with the full MS data.- x
A
MsBackendPyobject- columns
For
spectraData():characterwith the names of columns (spectra variables) to retrieve. Defaults tospectraVariables(object). ForpeaksData():characterwith the names of the peaks variables to retrieve.- drop
For
spectraData()andpeaksData():logical(1)whether, when a single column is requested, the data should be returned as avectorinstead of adata.frameormatrix.- value
Replacement value(s).
- name
For
$:character(1)with the name of the variable to retrieve.- backend
For
setBackend():MsBackendPyinstance.- applyProcessing
For
setBackend():logical(1)whether any cached data manipulation operations should be applied to the (peaks) data before transferring the data to Python. Defaults toapplyProcessing = TRUE.
Details
The MsBackendPy keeps only a reference to the MS data in Python (i.e. the
name of the variable in Python) as well as an index pointing to the
individual spectra in Python but no other data. Any data requested from
the MsBackendPy is accessed and translated on-the-fly from the Python
variable. The MsBackendPy is thus an interface to the Python MS data
structure, but not a data container. All changes to the MS data in the
Python variable (performed e.g. in Python) immediately affect any
MsBackendPy instances pointing to this variable.
All Subset operations on a MsBackendPy with e.g. [ are delayed
meaning that the data in Python is not immediately changed, but that
only the index to the individual spectrum objects in Python (which is stored
within the backend) is updated. Importantly, however, any replacement
operation such as $<- or rtime<- will realize the subset also in Python
(i.e., subset and change the order of the data in Python according to the
index in R). Be aware that this might cause data corruption if two
MsBackendPy instances use to the same data in Python (i.e., refer to the
same Python variable). See pyspec_copy_on_replace() for a strategy to
avoid such data corruption.
Special care must be taken if the MS data structure in Python is subset or
its order is changed (e.g. by another process). In that case it might be
needed to re-index the backend using the reindex() function:
object <- reindex(object). This will update (replace) the index to the
individual spectra in Python which is stored within the backend.
Note
As mentioned in the details section the MS data is completely stored in
Python and the backend only references to this data through the name of
the variable in Python. Thus, each time MS data is requested from the
backend, it is retrieved in its current state.
If for example data was transformed or metadata added or removed in the
Python object, it immediately affects the Spectra object in R.
Python metadata names vs spectra variables
The matchms library in Python supports arbitrary metadata assigned to
a spectrum, similar to Spectra's spectra variables concept. However,
matchms and Spectra use different names for the same metadata/variables.
To support this, MsBackendPy implements spectraVariableMapping which
allows to link spectra variable names to matchms metadata names. This
mapping can be provided with the spectraVariableMapping parameter to the
backendInitialize() function. Note that matchms only supports metadata
names in lower case or snake_case. Spectra variables are thus
automatically renamed and mapped to the lower case variants if needed. Also,
be aware that matchms automatically renames certain metadata fields:
a metadata "NAME" will be automatically renamed to "compound_name".
See also setSpectraVariableMapping() for more information.
MsBackendPy methods
The MsBackendPy supports all methods defined by the Spectra::MsBackend()
interface for access to MS data. Details on the invidual functions can also
be found in the main documentation in the Spectra package (i.e. for
Spectra::MsBackend()). Here we provide information for functions with
specific properties of the backend.
backendInitialize(): this method can be used to either initialize the backend with data from a referenced and **existing ** MS data structure in Python, or, through parameterdata, first convert and store the provided data to a Python MS data structure and then initialize the backend pointing to this referenced variable (Python attribute). In both cases, the name of the Python attribute needs to be provided with the parameterpythonVariableName. The mapping between the spectra variable names in R and the related Python metadata variables can be specified with thespectraVariableMappingparameter. It has to be a namedcharacterwith names being the spectra variables and the values the respective name for the metadata in the Python MS data structure. It defaults todefaultSpectraVariableMapping()which returns the mapping of some core spectra variables for the matchms Python library. Be aware that only those spectra variables specified with this parameter are mapped and translated between R and Python. ForbackendInitialize()with parameterdataprovided, only the variables defined byspectraVariableMapping, and available indata, are converted and stored in Python. Also note that, for efficiency reasons, core spectra variables (those listed bySpectra::coreSpectraVariables()) defined withspectraVariableMappingbut that have only missing values, are ignored. ParameterpythonLibrarymust be used to specify the Python library representing the MS data in Python. It can be eitherpythonLibrary = "matchms"(the default) orpythonLibrary = "spectrum_utils". The function returns an initialized instance ofMsBackendPy. See examples below for different settings and conversion of spectra variables.setBackend(): change the backend from aSpectraobject toMsBackendPy. The function internally usesbackendInitialize()(see above) to convert and store the relevant data to Python. By default, withapplyProcessing = TRUE, all data manipulation operations are applied to the data before storing them to Python. Note that only spectra variables defined by parameterspectraVariableMappingare transferred to Python.intensity(),intensity()<-: get or replace the intensity values.intensity()returns aNumericListof length equal to the number of spectra with each element being the intensity values of the individual mass peaks per spectrum.intensity()<-takes the same list-like structure as input parameter. Both the number of spectra and the number of peaks must match the length of the spectra and the number of existing mass peaks. To change the number of peaks use thepeaksData()<-method instead that replaces the m/z and intensity values at the same time.mz(),mz()<-: get or replace the m/z values.mz()returns aNumericListof length equal to the number of spectra with each element being the m/z values of the individual mass peaks per spectrum.mz()<-takes the same list-like structure as input parameter. Both the number of spectra and the number of peaks must match the length of the spectra and the number of existing mass peaks. To change the number of peaks use thepeaksData()<-method instead that replaces the m/z and intensity values at the same time.peaksData(): extracts the peaks data matrices from the backend. Python code is applied to the data structure in Python to extract the m/z and intensity values as a list of (numpy) arrays. These are then translated into an Rlistof two-columnnumericmatrices. Because Python does not allow to name columns of an array, an additional loop in R is required to set the column names to"mz"and"intensity".peaksData()<-: replaces the full peaks data (i.e., m/z and intensity values) for all spectra. Parametervaluehas to be alist-like structure with each element being anumericmatrix with one column (named"mz") containing the spectrum's m/z and one column (named"intensity") with the intensity values.spectraData(): extracts the spectra data from the backend. Which spectra variables are translated and retrieved from the Python objects depends on the backend'sspectraVariableMapping(). All metadata names defined are retrieved and added to the returnedDataFrame(with eventually missing core spectra variables filled withNA).spectraData()<-: replaces the full spectra (+ peaks) data of the backend with the values provided with the submittedDataFrame. The number of rows of thisDataFramehas to match the number of spectra ofobject(i.e., being equal tolength(object)) and theDataFramemust also contain the spectras' m/z and intensity values (in columns named"mz"and"intensity").spectraNames(),spectraNames()<-: extracts, respectively, adds (replaces) names for the individual spectra. These are stored as spectra variable/metadata field"spectrum_name"in the Python representation. This is only supported if the Python matchms library is used, the spectrum_utils library does not support arbitrary spectra names.spectraVariables(): retrieves available spectra variables, which include the names of all metadata attributes in thematchms.Spectrumobjects and the core spectra variablesSpectra::coreSpectraVariables().spectraVariableMapping(): get the currently defined mapping forspectraVariables()of the backend.spectraVariableMapping<-: replaces thespectraVariableMappingof the backend (seesetSpectraVariableMapping()for details and description of the expected format).$,$<-: extract or add/replace values for a spectra variable from/in the backend. If the spectrum_utils Python library is used, only a restricted set of spectra variables are supported, i.e.,precursorMz,precursorCharge,rtimeandscanIndex.
Additional helper and utility functions
reindex(): update the internal index to match1:length(object). This function is useful if the original data referenced by the backend was subset or re-ordered by a different process (or a function in Python).
See also
pyspec_copy_on_replace() to ensure MS data in Python gets copied
on any data replacement operation.
Examples
## Loading an example MGF file provided by the SpectriPy package.
## As an alternative, the data could also be imported directly in Python
## using:
## import matchms
## from matchms.importing import load_from_mgf
## s_p = list(load_from_mgf(r.fl))
library(Spectra)
#> Loading required package: S4Vectors
#> Loading required package: stats4
#> Loading required package: BiocGenerics
#> Loading required package: generics
#>
#> Attaching package: ‘generics’
#> The following objects are masked from ‘package:base’:
#>
#> as.difftime, as.factor, as.ordered, intersect, is.element, setdiff,
#> setequal, union
#>
#> Attaching package: ‘BiocGenerics’
#> The following objects are masked from ‘package:stats’:
#>
#> IQR, mad, sd, var, xtabs
#> The following objects are masked from ‘package:base’:
#>
#> Filter, Find, Map, Position, Reduce, anyDuplicated, aperm, append,
#> as.data.frame, basename, cbind, colnames, dirname, do.call,
#> duplicated, eval, evalq, get, grep, grepl, is.unsorted, lapply,
#> mapply, match, mget, order, paste, pmax, pmax.int, pmin, pmin.int,
#> rank, rbind, rownames, sapply, saveRDS, table, tapply, unique,
#> unsplit, which.max, which.min
#>
#> Attaching package: ‘S4Vectors’
#> The following object is masked from ‘package:utils’:
#>
#> findMatches
#> The following objects are masked from ‘package:base’:
#>
#> I, expand.grid, unname
#> Loading required package: BiocParallel
library(SpectriPy)
library(MsBackendMgf)
fl <- system.file("extdata", "mgf", "test.mgf", package = "SpectriPy")
s <- Spectra(fl, source = MsBackendMgf())
#> Start data import from 1 files ...
#> done
s
#> MSn data (Spectra) with 100 spectra in a MsBackendMgf backend:
#> msLevel rtime scanIndex
#> <integer> <numeric> <integer>
#> 1 2 NA NA
#> 2 2 NA NA
#> 3 2 NA NA
#> 4 2 NA NA
#> 5 2 NA NA
#> ... ... ... ...
#> 96 2 NA NA
#> 97 2 NA NA
#> 98 2 NA NA
#> 99 2 NA NA
#> 100 NA NA NA
#> ... 20 more variables/columns.
## Translating the MS data to Python and assigning it to a variable
## named "s_p" in the (*reticulate*'s) `py` Python environment. Assigning
## the variable to the Python environment has performance advantages, as
## any Python code applied to the MS data does not require any data
## conversions.
py_set_attr(py, "s_p", rspec_to_pyspec(s))
## Create a `MsBackendPy` representing an interface to the data in the
## "s_p" variable in Python:
be <- backendInitialize(MsBackendPy(), "s_p")
be
#> MsBackendPy with 100 spectra
#> Data stored in the "s_p" variable in Python
## Alternatively, by passing the full MS data with parameter `data`, the
## data is first converted to Python and the backend is initialized with
## that data. The `setBackend()` call from above internally uses this
## code to convert the data.
be <- backendInitialize(MsBackendPy(), "s_p3",
data = spectraData(s, c(spectraVariables(s), "mz", "intensity")))
## Create a Spectra object which this backend:
s_2 <- Spectra(be)
s_2
#> MSn data (Spectra) with 100 spectra in a MsBackendPy backend:
#> Data stored in the "s_p3" variable in Python
## An easier way to change the data representation of a `Spectra` object
## from R to Python is to use the `Spectra`'s `setBackend()` method
## selecting a `MsBackendPy` as the target backend representation:
s_2 <- setBackend(s, MsBackendPy(), pythonVariableName = "s_p2")
s_2
#> MSn data (Spectra) with 100 spectra in a MsBackendPy backend:
#> Data stored in the "s_p2" variable in Python
#> Processing:
#> Switch backend from MsBackendMgf to MsBackendPy [Tue May 12 06:49:50 2026]
## This moved the data from R to Python, storing it in a Python variable
## with the name `s_p2`. The resulting `s_2` is thus a `Spectra` object
## with all MS data however stored in Python.
## Note that by default only spectra variables that are part of
## `defaultSpectraVariableMapping()` are converted to Python
defaultSpectraVariableMapping()
#> precursorMz precursorIntensity
#> "precursor_mz" "precursor_intensity"
#> precursorCharge rtime
#> "charge" "retention_time"
#> collisionEnergy isolationWindowTargetMz
#> "collision_energy" "isolation_window_target_mz"
#> msLevel dataStorage
#> "ms_level" "data_storage"
## Thus, for example the precursor m/z is available in `s_2`, but other
## spectra variables from `s`, such as `"SMILES"` are not:
precursorMz(s)
#> [1] 259.0595 837.5318 250.1802 250.1802 250.1802 250.1802 250.1802 250.1802
#> [9] 195.0877 100.0757 154.0777 190.1352 175.1190 287.0574 83.0604 83.0604
#> [17] 195.0877 NA NA 780.5500 770.5700 179.1179 195.0877 195.0877
#> [25] 195.0877 425.1871 285.1809 224.0728 202.0854 161.0220 296.1412 239.1503
#> [33] 990.9768 239.0673 163.0314 303.1339 NA NA NA NA
#> [41] NA NA NA NA NA NA NA NA
#> [49] NA NA 296.1467 205.1911 317.2122 216.0000 207.0000 125.0000
#> [57] 164.0000 256.0000 138.0000 256.0000 436.9460 866.6100 254.2478 335.0512
#> [65] 257.1285 239.0815 239.0815 239.0815 301.1445 301.1445 302.3054 175.1190
#> [73] 175.1190 NA 497.3272 175.1189 175.1189 230.0958 350.1598 NA
#> [81] NA 611.1612 273.1386 385.2122 449.1078 447.0933 591.1719 339.2067
#> [89] 878.5001 NA 102.0600 465.4100 459.2748 NA 301.1445 301.1445
#> [97] 407.1347 304.1918 361.2020 NA
precursorMz(s_2)
#> [1] 259.0595 837.5318 250.1802 250.1802 250.1802 250.1802 250.1802 250.1802
#> [9] 195.0877 100.0757 154.0777 190.1352 175.1190 287.0574 83.0604 83.0604
#> [17] 195.0877 NA NA 780.5500 770.5700 179.1179 195.0877 195.0877
#> [25] 195.0877 425.1871 285.1809 224.0728 202.0854 161.0220 296.1412 239.1503
#> [33] 990.9768 239.0673 163.0314 303.1339 NA NA NA NA
#> [41] NA NA NA NA NA NA NA NA
#> [49] NA NA 296.1467 205.1911 317.2122 216.0000 207.0000 125.0000
#> [57] 164.0000 256.0000 138.0000 256.0000 436.9460 866.6100 254.2478 335.0512
#> [65] 257.1285 239.0815 239.0815 239.0815 301.1445 301.1445 302.3054 175.1190
#> [73] 175.1190 NA 497.3272 175.1189 175.1189 230.0958 350.1598 NA
#> [81] NA 611.1612 273.1386 385.2122 449.1078 447.0933 591.1719 339.2067
#> [89] 878.5001 NA 102.0600 465.4100 459.2748 NA 301.1445 301.1445
#> [97] 407.1347 304.1918 361.2020 NA
s$SMILES |> head()
#> [1] "CC1=CC(=CC2=C1C3=CC(=CC(=C3C(=O)O2)O)O)O"
#> [2] "CC[C@@H]1[C@@]([C@@H]([C@H](/C(=N/OCOCCOC)/[C@@H](C[C@@]([C@@H]([C@H]([C@@H]([C@H](C(=O)O1)C)O[C@H]2C[C@@]([C@H]([C@@H](O2)C)O)(C)OC)C)O[C@H]3[C@@H]([C@H](C[C@H](O3)C)N(C)C)O)(C)O)C)C)O)(C)O"
#> [3] "CN(C)C[C@@H]1CCCC([C@H]1C2=CC(=CC=C2)O)O"
#> [4] "CN(C)C[C@@H]1CCCC([C@H]1C2=CC(=CC=C2)O)O"
#> [5] "CN(C)C[C@H]1CCCC[C@@]1(C2=CC(=CC=C2)O)O"
#> [6] "CN(C)C[C@H]1CCCC[C@@]1(C2=CC(=CC=C2)O)O"
## s_2$SMILES would throw an error.
## To also translate this spectra variable, it needs to be included and
## specified with the `spectraVariableMapping` parameter. The easiest
## approach is to use the `spectraVariableMapping()` function adding in
## addition to the default mapping for the Python library (`"matchms"`)
## also the mapping of additional spectra variables that should be converted:
s_2 <- setBackend(s, MsBackendPy(), pythonVariableName = "s_p2",
spectraVariableMapping = spectraVariableMapping("matchms", c(SMILES = "smiles")))
s_2$SMILES |> head()
#> [1] "CC1=CC(=CC2=C1C3=CC(=CC(=C3C(=O)O2)O)O)O"
#> [2] "CC[C@@H]1[C@@]([C@@H]([C@H](/C(=N/OCOCCOC)/[C@@H](C[C@@]([C@@H]([C@H]([C@@H]([C@H](C(=O)O1)C)O[C@H]2C[C@@]([C@H]([C@@H](O2)C)O)(C)OC)C)O[C@H]3[C@@H]([C@H](C[C@H](O3)C)N(C)C)O)(C)O)C)C)O)(C)O"
#> [3] "CN(C)C[C@@H]1CCCC([C@H]1C2=CC(=CC=C2)O)O"
#> [4] "CN(C)C[C@@H]1CCCC([C@H]1C2=CC(=CC=C2)O)O"
#> [5] "CN(C)C[C@H]1CCCC[C@@]1(C2=CC(=CC=C2)O)O"
#> [6] "CN(C)C[C@H]1CCCC[C@@]1(C2=CC(=CC=C2)O)O"
## Available spectra variables: these include, next to the *core* spectra
## variables, also the names of all metadata stored in the `matchms.Spectrum`
## objects.
spectraVariables(s_2)
#> [1] "msLevel" "rtime"
#> [3] "acquisitionNum" "scanIndex"
#> [5] "dataStorage" "dataOrigin"
#> [7] "centroided" "smoothed"
#> [9] "polarity" "precScanNum"
#> [11] "precursorMz" "precursorIntensity"
#> [13] "precursorCharge" "collisionEnergy"
#> [15] "isolationWindowLowerMz" "isolationWindowTargetMz"
#> [17] "isolationWindowUpperMz" "SMILES"
## Get the full peaks data:
peaksData(s_2)
#> List of length 100
## Get the peaks from the first spectrum
peaksData(s_2)[[1L]]
#> mz intensity
#> [1,] 213.0546 754969.6
#> [2,] 241.0495 1058878.8
#> [3,] 259.0601 20211204.0
## Get the full spectra data:
spectraData(s_2)
#> DataFrame with 100 rows and 18 columns
#> msLevel rtime acquisitionNum scanIndex dataStorage dataOrigin
#> <integer> <numeric> <integer> <integer> <character> <character>
#> 1 2 NA NA NA s_p2 NA
#> 2 2 NA NA NA s_p2 NA
#> 3 2 NA NA NA s_p2 NA
#> 4 2 NA NA NA s_p2 NA
#> 5 2 NA NA NA s_p2 NA
#> ... ... ... ... ... ... ...
#> 96 2 NA NA NA s_p2 NA
#> 97 2 NA NA NA s_p2 NA
#> 98 2 NA NA NA s_p2 NA
#> 99 2 NA NA NA s_p2 NA
#> 100 NA NA NA NA s_p2 NA
#> centroided smoothed polarity precScanNum precursorMz precursorIntensity
#> <logical> <logical> <integer> <integer> <numeric> <numeric>
#> 1 NA NA NA NA 259.060 NA
#> 2 NA NA NA NA 837.532 NA
#> 3 NA NA NA NA 250.180 NA
#> 4 NA NA NA NA 250.180 NA
#> 5 NA NA NA NA 250.180 NA
#> ... ... ... ... ... ... ...
#> 96 NA NA NA NA 301.144 NA
#> 97 NA NA NA NA 407.135 NA
#> 98 NA NA NA NA 304.192 NA
#> 99 NA NA NA NA 361.202 NA
#> 100 NA NA NA NA NA NA
#> precursorCharge collisionEnergy isolationWindowLowerMz
#> <integer> <numeric> <numeric>
#> 1 1 NA NA
#> 2 1 NA NA
#> 3 1 NA NA
#> 4 1 NA NA
#> 5 1 NA NA
#> ... ... ... ...
#> 96 1 NA NA
#> 97 1 NA NA
#> 98 1 NA NA
#> 99 1 NA NA
#> 100 1 NA NA
#> isolationWindowTargetMz isolationWindowUpperMz SMILES
#> <numeric> <numeric> <character>
#> 1 NA NA CC1=CC(=CC2=C1C3=CC(..
#> 2 NA NA CC[C@@H]1[C@@]([C@@H..
#> 3 NA NA CN(C)C[C@@H]1CCCC([C..
#> 4 NA NA CN(C)C[C@@H]1CCCC([C..
#> 5 NA NA CN(C)C[C@H]1CCCC[C@@..
#> ... ... ... ...
#> 96 NA NA CC(CC1=CC(=C(C=C1)O)..
#> 97 NA NA CC1C(=O)OC2C1(C34C(=..
#> 98 NA NA CC(C)/C=C/CCCCC(=O)N..
#> 99 NA NA C[C@]12CCC(=O)C=C1CC..
#> 100 NA NA c(c3)ccc(c3)CC(=O)NC..
## Get the m/z values
mz(s_2)
#> NumericList of length 100
#> [[1]] 213.0546 241.0495 259.0601
#> [[2]] 116.0712 116.1074 127.0759 142.1231 ... 398.2568 679.4443 680.4436
#> [[3]] 232.1689 250.18 251.1831 252.1852
#> [[4]] 145.0638 187.1102 232.1687 233.1721 250.1795 251.1824 252.1862
#> [[5]] 232.1689 250.18 251.1831 252.1852
#> [[6]] 145.0638 187.1102 232.1687 233.1721 250.1795 251.1824 252.1862
#> [[7]] 232.1689 250.18 251.1831 252.1852
#> [[8]] 145.0638 187.1102 232.1687 233.1721 250.1795 251.1824 252.1862
#> [[9]] 135.0432 138.0632 163.0375 195.088
#> [[10]] 55.0537 56.0487 58.0281 58.0644 ... 71.0493 72.0792 82.0652 100.0763
#> ...
#> <90 more elements>
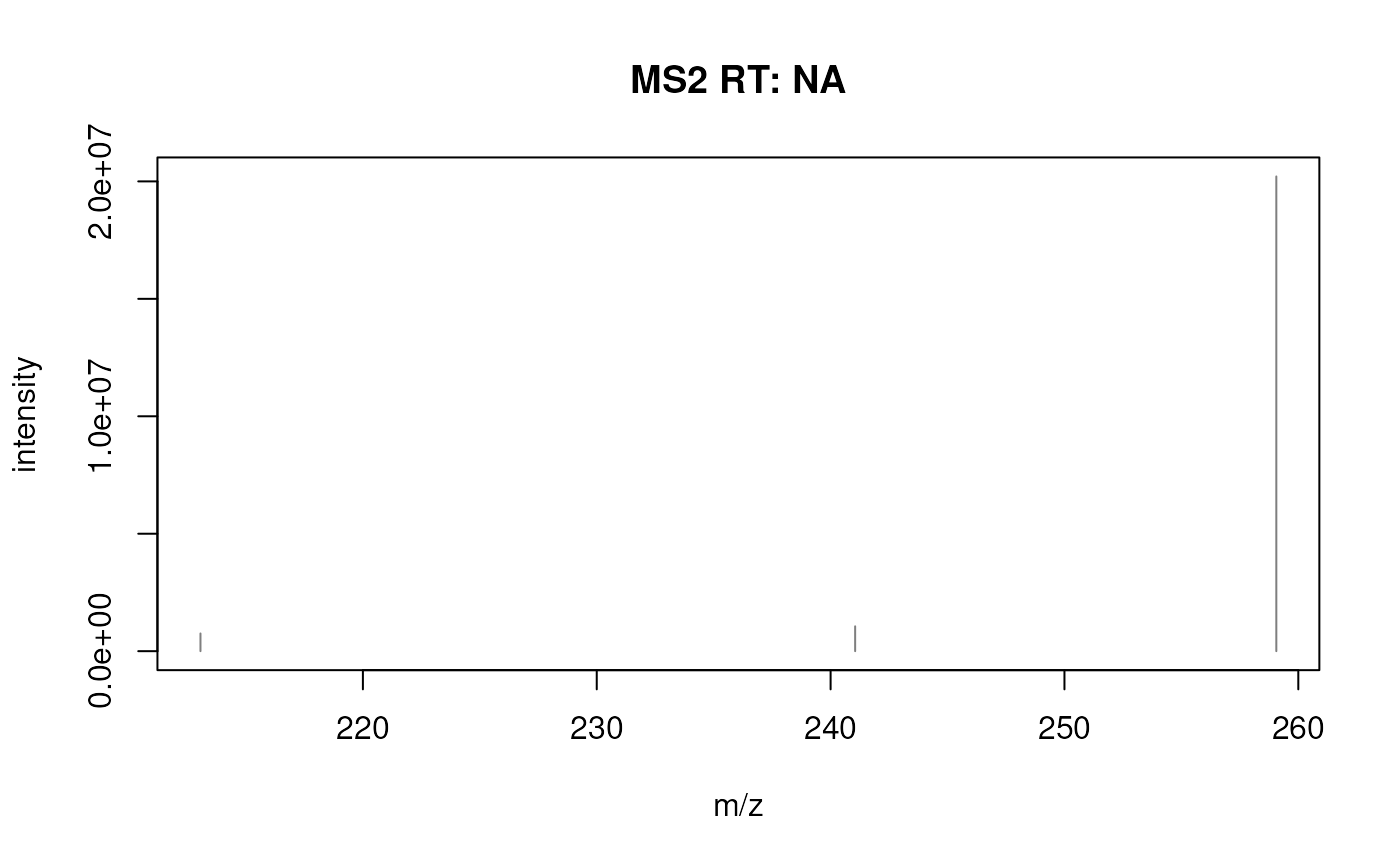
## Plot the first spectrum
plotSpectra(s_2[1L])
 ########
## Using the spectrum_utils Python library
## Below we convert the data to a list of `MsmsSpectrum` object from the
## spectrum_utils library.
py_set_attr(py, "su_p", rspec_to_pyspec(s,
spectraVariableMapping("spectrum_utils"), "spectrum_utils"))
## Create a MsBackendPy representing this data. Importantly, we need to
## specify the Python library using the `pythonLibrary` parameter and
## ideally also set the `spectraVariableMapping` to the one specific for
## that library.
be <- backendInitialize(MsBackendPy(), "su_p",
spectraVariableMapping = spectraVariableMapping("spectrum_utils"),
pythonLibrary = "spectrum_utils")
be
#> MsBackendPy with 100 spectra
#> Data stored in the "su_p" variable in Python
## Get the peaks data for the first 3 spectra
peaksData(be[1:3])
#> [[1]]
#> mz intensity
#> [1,] 213.0546 754969.6
#> [2,] 241.0495 1058878.8
#> [3,] 259.0601 20211204.0
#>
#> [[2]]
#> mz intensity
#> [1,] 116.0712 15660
#> [2,] 116.1074 28164
#> [3,] 127.0759 8616
#> [4,] 142.1231 4516
#> [5,] 158.1184 855092
#> [6,] 159.1218 68248
#> [7,] 380.2469 6148
#> [8,] 398.2568 8996
#> [9,] 679.4443 10596
#> [10,] 680.4436 5176
#>
#> [[3]]
#> mz intensity
#> [1,] 232.1689 20176
#> [2,] 250.1800 2073248
#> [3,] 251.1831 282228
#> [4,] 252.1852 21640
#>
## Get the full spectraData
spectraData(be)
#> Warning: NAs introduced by coercion
#> DataFrame with 100 rows and 19 columns
#> msLevel rtime acquisitionNum scanIndex mz
#> <integer> <numeric> <integer> <integer> <NumericList>
#> 1 NA NaN NA NA 213.055,241.049,259.060
#> 2 NA NaN NA NA 116.071,116.107,127.076,...
#> 3 NA NaN NA NA 232.169,250.180,251.183,...
#> 4 NA NaN NA NA 145.064,187.110,232.169,...
#> 5 NA NaN NA NA 232.169,250.180,251.183,...
#> ... ... ... ... ... ...
#> 96 NA NaN NA NA 108.023,109.028,122.037,...
#> 97 NA NaN NA NA 277.099,289.147,291.149,...
#> 98 NA NaN NA NA 168.139,168.177,168.230,...
#> 99 NA NaN NA NA 106.764,116.930,125.058,...
#> 100 NA NaN NA NA 100,101,122,...
#> intensity dataStorage dataOrigin centroided smoothed
#> <NumericList> <character> <character> <logical> <logical>
#> 1 754970, 1058879,20211204 su_p NA NA NA
#> 2 15660,28164, 8616,... su_p NA NA NA
#> 3 20176,2073248, 282228,... su_p NA NA NA
#> 4 864, 2548,10716,... su_p NA NA NA
#> 5 20176,2073248, 282228,... su_p NA NA NA
#> ... ... ... ... ... ...
#> 96 224, 295,1801,... su_p NA NA NA
#> 97 21,26,27,... su_p NA NA NA
#> 98 24842, 882, 182,... su_p NA NA NA
#> 99 39, 51,108,... su_p NA NA NA
#> 100 47,153, 39,... su_p NA NA NA
#> polarity precScanNum precursorMz precursorIntensity precursorCharge
#> <integer> <integer> <numeric> <numeric> <integer>
#> 1 NA NA 259.060 NA 1
#> 2 NA NA 837.532 NA 1
#> 3 NA NA 250.180 NA 1
#> 4 NA NA 250.180 NA 1
#> 5 NA NA 250.180 NA 1
#> ... ... ... ... ... ...
#> 96 NA NA 301.144 NA 1
#> 97 NA NA 407.135 NA 1
#> 98 NA NA 304.192 NA 1
#> 99 NA NA 361.202 NA 1
#> 100 NA NA NA NA 1
#> collisionEnergy isolationWindowLowerMz isolationWindowTargetMz
#> <numeric> <numeric> <numeric>
#> 1 NA NA NA
#> 2 NA NA NA
#> 3 NA NA NA
#> 4 NA NA NA
#> 5 NA NA NA
#> ... ... ... ...
#> 96 NA NA NA
#> 97 NA NA NA
#> 98 NA NA NA
#> 99 NA NA NA
#> 100 NA NA NA
#> isolationWindowUpperMz
#> <numeric>
#> 1 NA
#> 2 NA
#> 3 NA
#> 4 NA
#> 5 NA
#> ... ...
#> 96 NA
#> 97 NA
#> 98 NA
#> 99 NA
#> 100 NA
## Extract the precursor m/z
be$precursorMz
#> [1] 259.0595 837.5318 250.1802 250.1802 250.1802 250.1802 250.1802 250.1802
#> [9] 195.0877 100.0757 154.0777 190.1352 175.1190 287.0574 83.0604 83.0604
#> [17] 195.0877 NA NA 780.5500 770.5700 179.1179 195.0877 195.0877
#> [25] 195.0877 425.1871 285.1809 224.0728 202.0854 161.0220 296.1412 239.1503
#> [33] 990.9768 239.0673 163.0314 303.1339 NA NA NA NA
#> [41] NA NA NA NA NA NA NA NA
#> [49] NA NA 296.1467 205.1911 317.2122 216.0000 207.0000 125.0000
#> [57] 164.0000 256.0000 138.0000 256.0000 436.9460 866.6100 254.2478 335.0512
#> [65] 257.1285 239.0815 239.0815 239.0815 301.1445 301.1445 302.3054 175.1190
#> [73] 175.1190 NA 497.3272 175.1189 175.1189 230.0958 350.1598 NA
#> [81] NA 611.1612 273.1386 385.2122 449.1078 447.0933 591.1719 339.2067
#> [89] 878.5001 NA 102.0600 465.4100 459.2748 NA 301.1445 301.1445
#> [97] 407.1347 304.1918 361.2020 NA
########
## Using the spectrum_utils Python library
## Below we convert the data to a list of `MsmsSpectrum` object from the
## spectrum_utils library.
py_set_attr(py, "su_p", rspec_to_pyspec(s,
spectraVariableMapping("spectrum_utils"), "spectrum_utils"))
## Create a MsBackendPy representing this data. Importantly, we need to
## specify the Python library using the `pythonLibrary` parameter and
## ideally also set the `spectraVariableMapping` to the one specific for
## that library.
be <- backendInitialize(MsBackendPy(), "su_p",
spectraVariableMapping = spectraVariableMapping("spectrum_utils"),
pythonLibrary = "spectrum_utils")
be
#> MsBackendPy with 100 spectra
#> Data stored in the "su_p" variable in Python
## Get the peaks data for the first 3 spectra
peaksData(be[1:3])
#> [[1]]
#> mz intensity
#> [1,] 213.0546 754969.6
#> [2,] 241.0495 1058878.8
#> [3,] 259.0601 20211204.0
#>
#> [[2]]
#> mz intensity
#> [1,] 116.0712 15660
#> [2,] 116.1074 28164
#> [3,] 127.0759 8616
#> [4,] 142.1231 4516
#> [5,] 158.1184 855092
#> [6,] 159.1218 68248
#> [7,] 380.2469 6148
#> [8,] 398.2568 8996
#> [9,] 679.4443 10596
#> [10,] 680.4436 5176
#>
#> [[3]]
#> mz intensity
#> [1,] 232.1689 20176
#> [2,] 250.1800 2073248
#> [3,] 251.1831 282228
#> [4,] 252.1852 21640
#>
## Get the full spectraData
spectraData(be)
#> Warning: NAs introduced by coercion
#> DataFrame with 100 rows and 19 columns
#> msLevel rtime acquisitionNum scanIndex mz
#> <integer> <numeric> <integer> <integer> <NumericList>
#> 1 NA NaN NA NA 213.055,241.049,259.060
#> 2 NA NaN NA NA 116.071,116.107,127.076,...
#> 3 NA NaN NA NA 232.169,250.180,251.183,...
#> 4 NA NaN NA NA 145.064,187.110,232.169,...
#> 5 NA NaN NA NA 232.169,250.180,251.183,...
#> ... ... ... ... ... ...
#> 96 NA NaN NA NA 108.023,109.028,122.037,...
#> 97 NA NaN NA NA 277.099,289.147,291.149,...
#> 98 NA NaN NA NA 168.139,168.177,168.230,...
#> 99 NA NaN NA NA 106.764,116.930,125.058,...
#> 100 NA NaN NA NA 100,101,122,...
#> intensity dataStorage dataOrigin centroided smoothed
#> <NumericList> <character> <character> <logical> <logical>
#> 1 754970, 1058879,20211204 su_p NA NA NA
#> 2 15660,28164, 8616,... su_p NA NA NA
#> 3 20176,2073248, 282228,... su_p NA NA NA
#> 4 864, 2548,10716,... su_p NA NA NA
#> 5 20176,2073248, 282228,... su_p NA NA NA
#> ... ... ... ... ... ...
#> 96 224, 295,1801,... su_p NA NA NA
#> 97 21,26,27,... su_p NA NA NA
#> 98 24842, 882, 182,... su_p NA NA NA
#> 99 39, 51,108,... su_p NA NA NA
#> 100 47,153, 39,... su_p NA NA NA
#> polarity precScanNum precursorMz precursorIntensity precursorCharge
#> <integer> <integer> <numeric> <numeric> <integer>
#> 1 NA NA 259.060 NA 1
#> 2 NA NA 837.532 NA 1
#> 3 NA NA 250.180 NA 1
#> 4 NA NA 250.180 NA 1
#> 5 NA NA 250.180 NA 1
#> ... ... ... ... ... ...
#> 96 NA NA 301.144 NA 1
#> 97 NA NA 407.135 NA 1
#> 98 NA NA 304.192 NA 1
#> 99 NA NA 361.202 NA 1
#> 100 NA NA NA NA 1
#> collisionEnergy isolationWindowLowerMz isolationWindowTargetMz
#> <numeric> <numeric> <numeric>
#> 1 NA NA NA
#> 2 NA NA NA
#> 3 NA NA NA
#> 4 NA NA NA
#> 5 NA NA NA
#> ... ... ... ...
#> 96 NA NA NA
#> 97 NA NA NA
#> 98 NA NA NA
#> 99 NA NA NA
#> 100 NA NA NA
#> isolationWindowUpperMz
#> <numeric>
#> 1 NA
#> 2 NA
#> 3 NA
#> 4 NA
#> 5 NA
#> ... ...
#> 96 NA
#> 97 NA
#> 98 NA
#> 99 NA
#> 100 NA
## Extract the precursor m/z
be$precursorMz
#> [1] 259.0595 837.5318 250.1802 250.1802 250.1802 250.1802 250.1802 250.1802
#> [9] 195.0877 100.0757 154.0777 190.1352 175.1190 287.0574 83.0604 83.0604
#> [17] 195.0877 NA NA 780.5500 770.5700 179.1179 195.0877 195.0877
#> [25] 195.0877 425.1871 285.1809 224.0728 202.0854 161.0220 296.1412 239.1503
#> [33] 990.9768 239.0673 163.0314 303.1339 NA NA NA NA
#> [41] NA NA NA NA NA NA NA NA
#> [49] NA NA 296.1467 205.1911 317.2122 216.0000 207.0000 125.0000
#> [57] 164.0000 256.0000 138.0000 256.0000 436.9460 866.6100 254.2478 335.0512
#> [65] 257.1285 239.0815 239.0815 239.0815 301.1445 301.1445 302.3054 175.1190
#> [73] 175.1190 NA 497.3272 175.1189 175.1189 230.0958 350.1598 NA
#> [81] NA 611.1612 273.1386 385.2122 449.1078 447.0933 591.1719 339.2067
#> [89] 878.5001 NA 102.0600 465.4100 459.2748 NA 301.1445 301.1445
#> [97] 407.1347 304.1918 361.2020 NA
